Covary protocol portfolio is now available.
Machine learning approach to multi-locus Y-chromosome STR sequence profiling using Covary
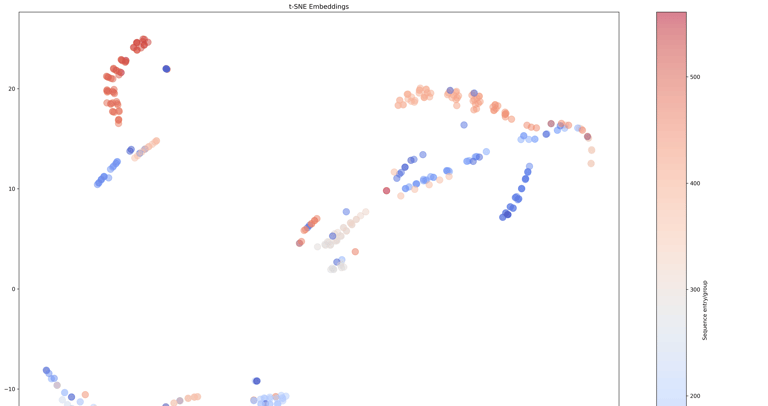
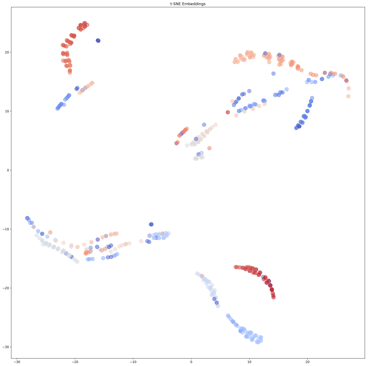
This protocol describes the use of Covary for rapid, alignment-free analysis of short tandem repeat (STR) sequence variation using Y-chromosomal STR loci from the NCBI STRSeq BioProject (PRJNA380347). The workflow demonstrates the ability of Covary to (i) compare STR sequence similarity directly from raw sequence data, (ii) perform multi-locus STR analysis in a single run without locus-wise concatenation, and (iii) resolve locus-specific and inter-locus relationships using machine learning-derived vector representations. Traditional forensic STR analysis relies on length-based allele designation and locus-by-locus interpretation. In contrast, this protocol illustrates a sequence-level, machine learning approach that captures internal repeat structure, flanking variation, and compositional features across multiple STR loci simultaneously. Covary enables scalable STR sequence comparison without manual alignment, custom scripting, or local software installation. This protocol is optimized for execution in Google Colab and may be adapted for applications in forensic genomics, population genetics, and STR database exploration.
12/26/2025
Covary is powered by:




© 2025 Marvin De los Santos
All rights reserved

Privacy | Terms | License Notice | Request Access | Procurement
Part of ChordexBio – algorithms empowering lives