Covary protocol portfolio is now available.
Machine learning-based phylogenetic analysis using Covary
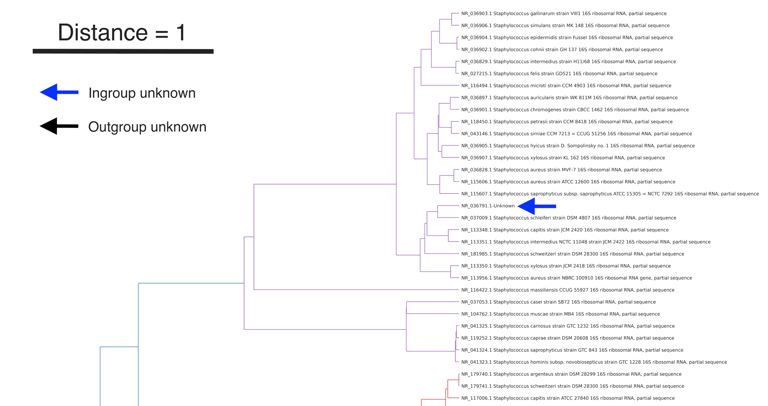
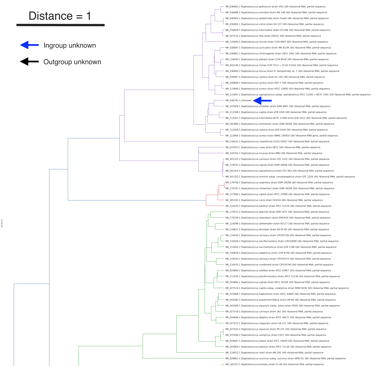
This protocol describes the operational workflow of Covary, a machine learning-based framework for large-scale phylogenetic analysis and species identification. This protocol enables users to perform phylogenetic inference directly from sequence data without requiring coding experience or local software installation. The workflow is applicable to v2.1 of Covary and accepts multi-FASTA sequence files as input (or training data) and provides configurable parameters for encoding, neural network inference, and downstream analysis. Covary is designed to scale to thousands of sequences and produces interoperable outputs compatible with Matplotlib and other python-based libraries, R, and other downstream visualization and statistical tools. The protocol is optimized for execution in a Google Colab environment, eliminating software maintenance and platform dependency. This protocol focuses on the methodical operations of Covary for tree reconstruction and species identification. Covary is available at https://github.com/mahvin92/Covary.
12/24/2025
Covary is powered by:




© 2025 Marvin De los Santos
All rights reserved

Privacy | Terms | License Notice | Request Access | Procurement
Part of ChordexBio – algorithms empowering lives