Covary protocol portfolio is now available.
Rapid, large-scale and multi-species phylogenomic analysis using Covary
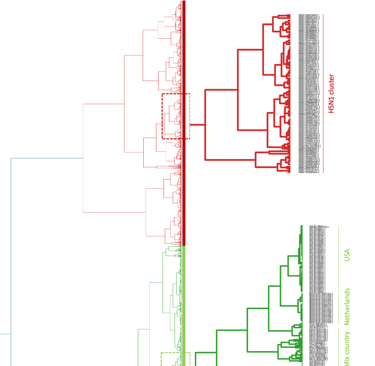
This protocol describes the use of Covary for fast, large-scale, multi-species phylogenomic analysis using complete genome sequences. The workflow is optimized for comparative phylogenomic inference across diverse taxa and is demonstrated using datasets associated with genomes of outbreak-causing viruses (SARS-CoV-2, dengue virus, measles virus, and alphainfluenza virus). Covary enables alignment-free phylogenomic analysis by encoding genomic sequences into translation-aware vector representations and applying machine learning–based similarity inference. The protocol supports thousands-scale datasets, requires no coding experience, and is designed to run in a Google Colab environment without local software installation or maintenance. This protocol focuses on phylogenomic-scale analysis across multiple viral species. Covary is accessible at https://covary.chordexbio.com/ or on GitHub at https://github.com/mahvin92/Covary.
12/26/2025
Covary is powered by:




© 2025 Marvin De los Santos
All rights reserved

Privacy | Terms | License Notice | Request Access | Procurement
Part of ChordexBio – algorithms empowering lives