Covary performs taxonomic classification similarly with alignment-based algorithms.
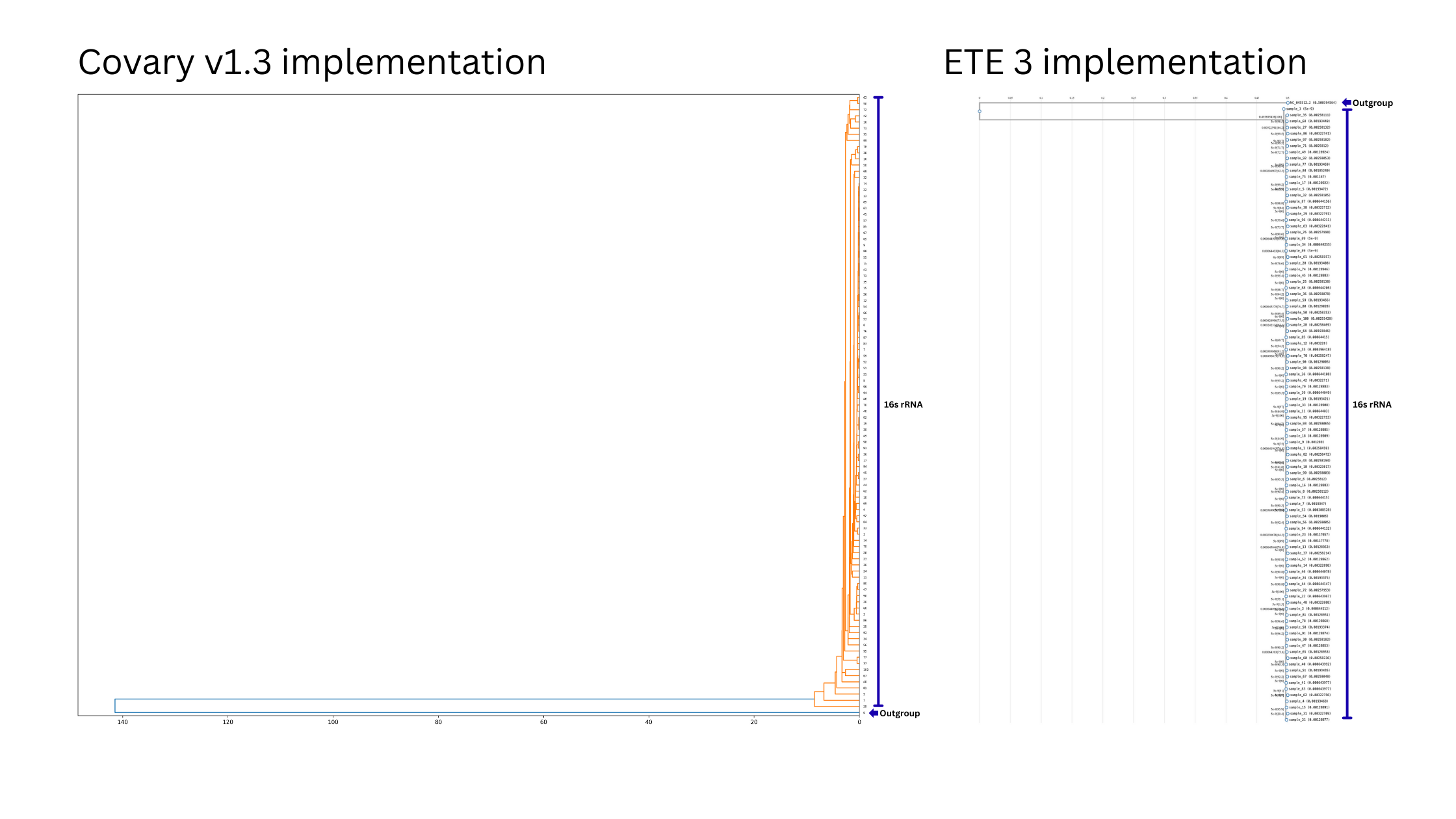
Covary has similar efficiency as conventional pipelines in sequence identification.
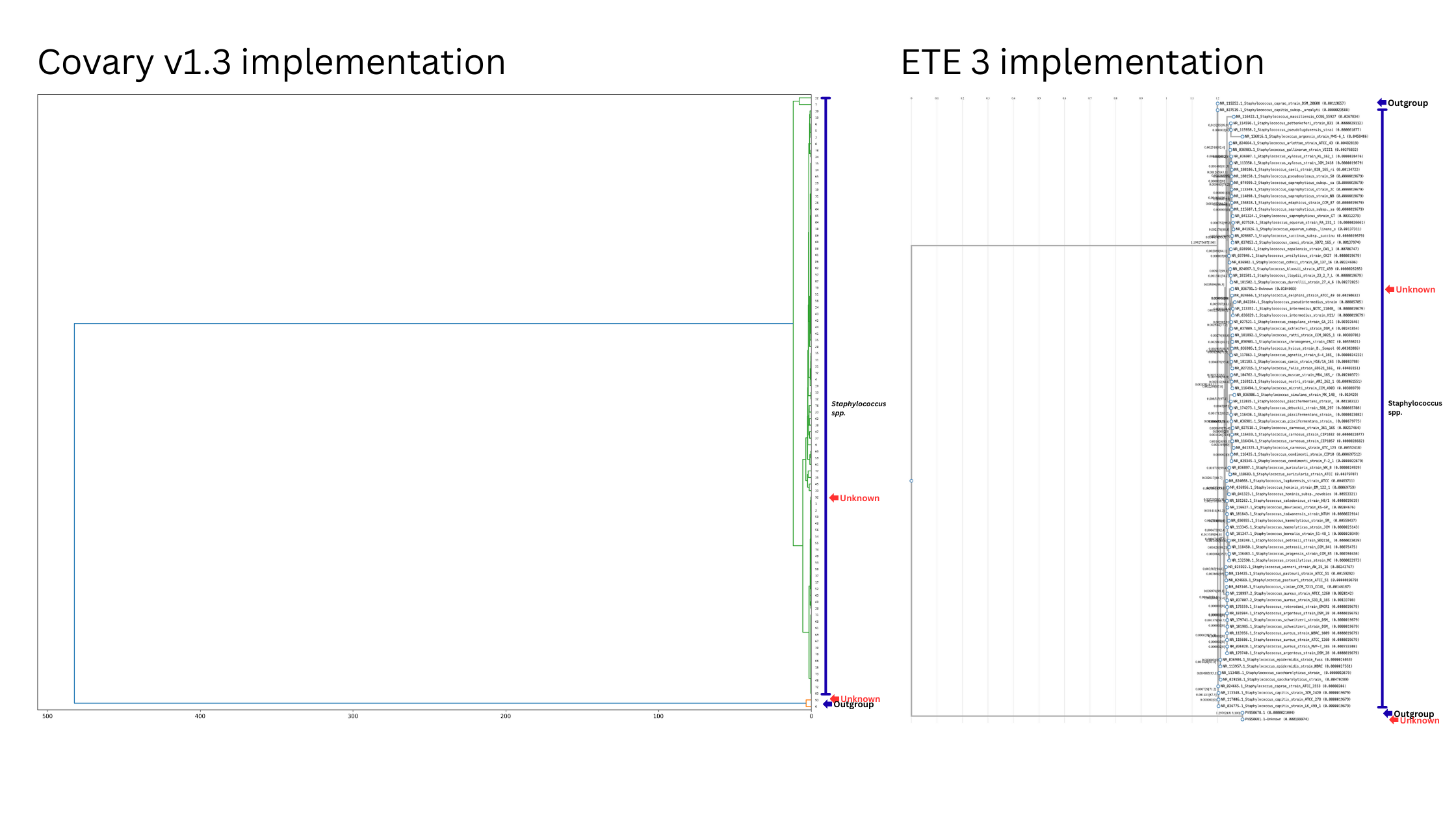
Covary outperforms conventional pipelines in providing sequence relationships.
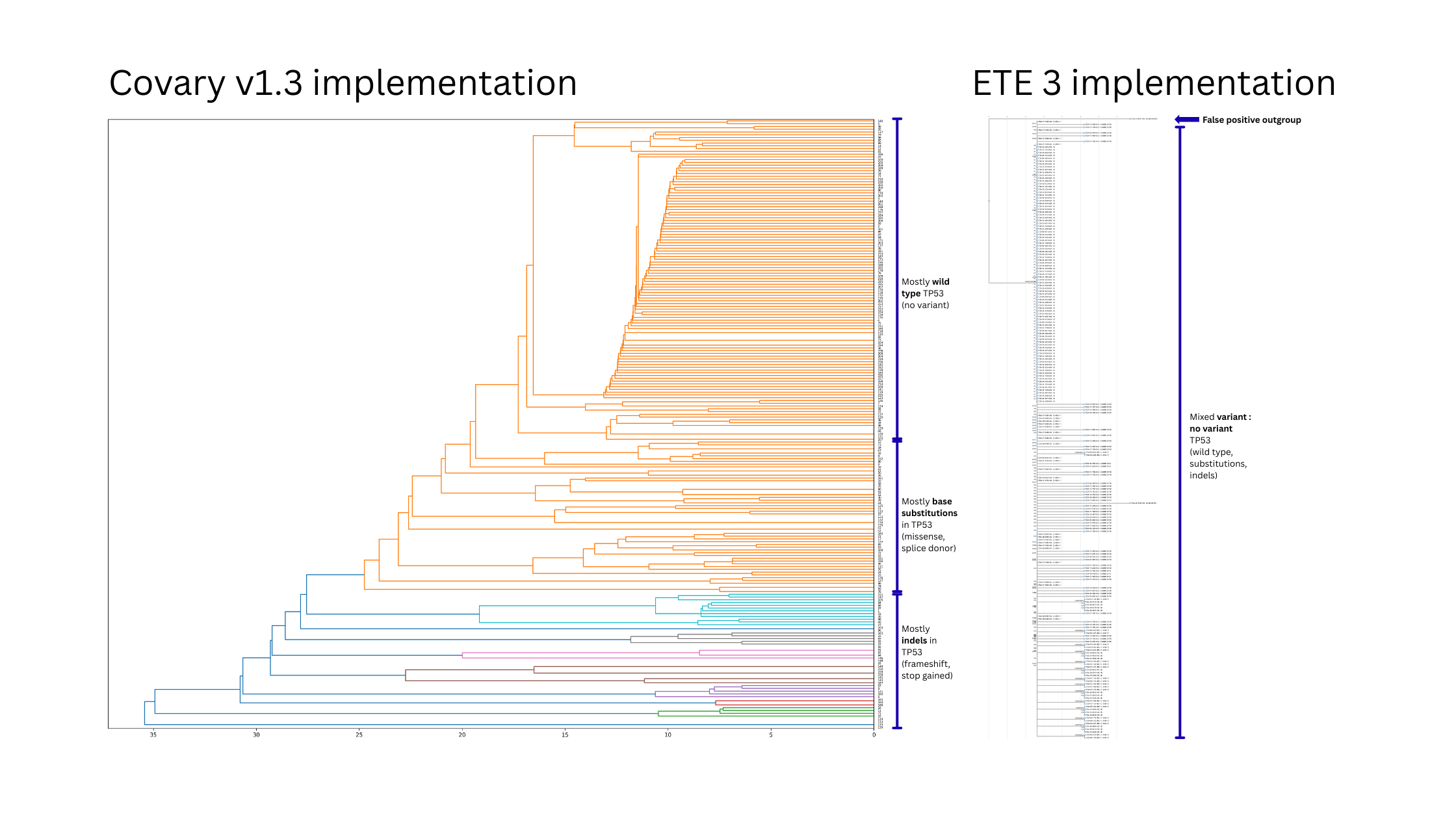
Covary features bring superior predictive context not found in conventional phylogenetics.
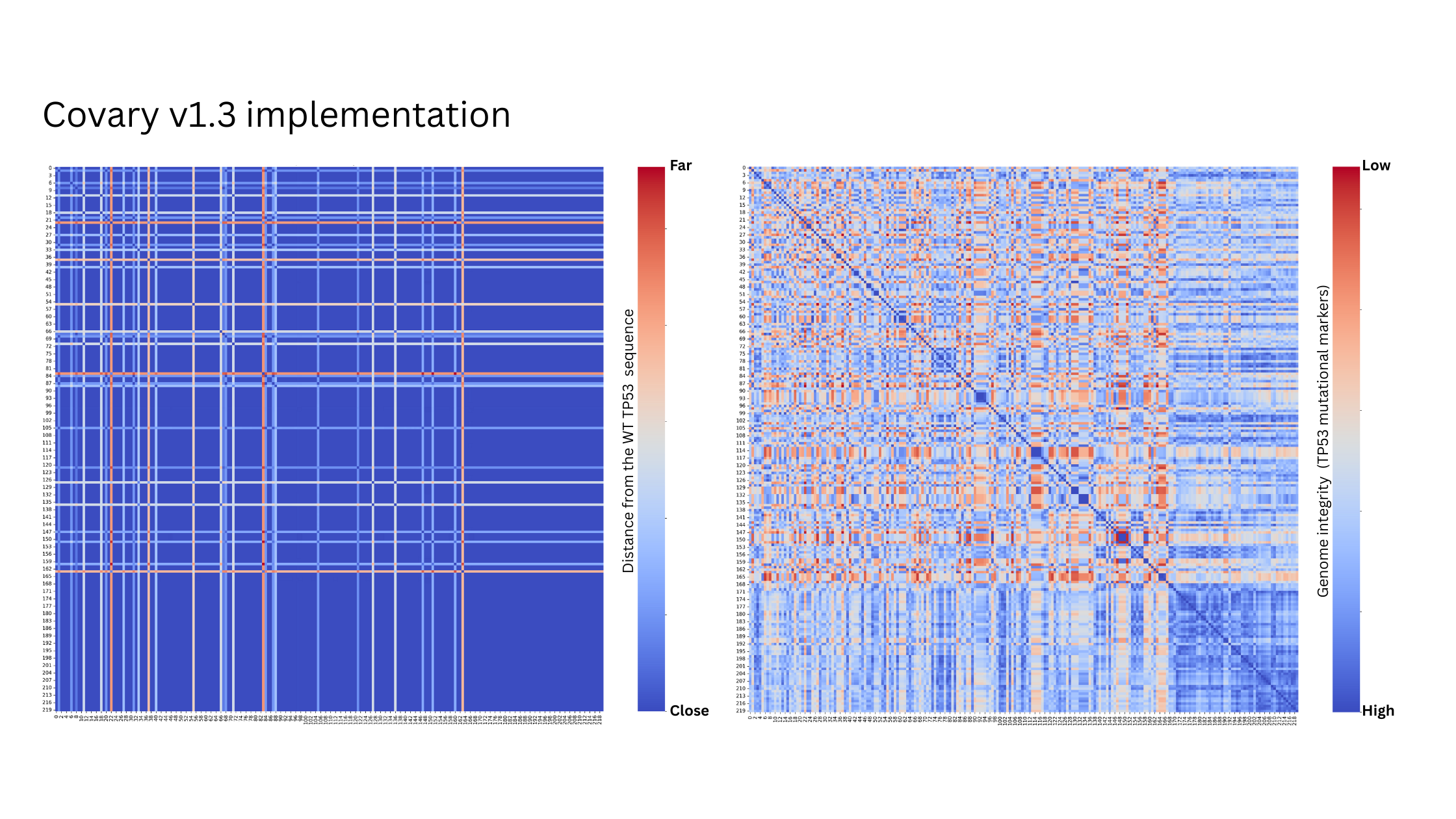
Comparison with the gold standard
Covary vs. alignment-based phylogeneticsCovary occupies a distinct computational niche from established tools such as MEGA, IQ-TREE, RAxML, and FastTree. The table below summarizes key differences to help researchers understand where Covary adds value and where traditional methods remain appropriate.
| Criterion | Gold standard (MEGA / IQ-TREE / RAxML) | Covary |
|---|---|---|
| Alignment requirement | Required — MSA is a prerequisite | None — alignment-free by design |
| Substitution model | Explicit evolutionary model (GTR, HKY, WAG, etc.) | No model assumption — distance from embedding space |
| Scale | Hundreds to low thousands (MSA bottleneck) | Hundreds to thousands; megascale in v3.0.1 beta |
| Speed | Slow at scale — MSA and model optimization rate-limiting | Fast — no MSA; k-mer encoding is lightweight |
| Translation awareness | Optional (codon models in some tools) | Built-in — codon-bound, relative-proximity encoding |
| Output | Newick trees, bootstrap values, model parameters | PCA/t-SNE/UMAP plots, heatmaps, dendrograms |
| Statistical support | Bootstrap, UFBoot, SH-aLRT available | Not currently implemented — future roadmap |
| Best suited for | High-precision species tree inference, publication-ready phylogenies | Rapid exploratory analysis, large-scale screening |
| Standardization | Widely peer-reviewed across thousands of studies | Experimental; validated in preprint studies — ongoing |
How to read this: Covary is not a direct replacement for MEGA, IQ-TREE, or RAxML — it solves a different problem. For high-confidence species trees with statistical support and publication-grade phylogeny reconstruction, alignment-based tools remain the gold standard. Covary's strength is speed and scale at the exploratory stage — rapidly screening large, heterogeneous datasets and generating embedding-based cluster hypotheses that can then be followed up with targeted alignment-based analyses on subsets of interest.